Pictures of the Day 2-27-08
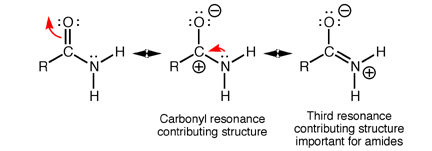
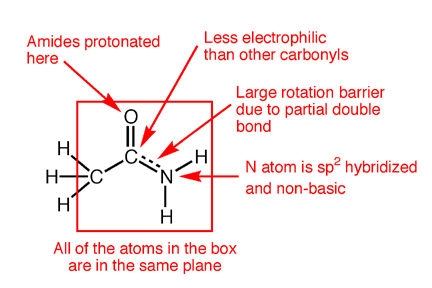
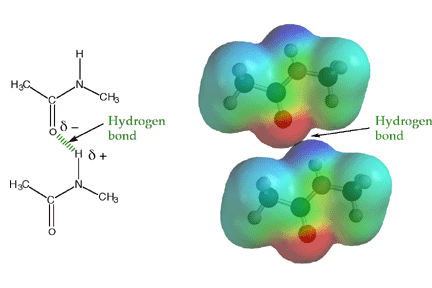
Amides have structural characteristics that are unique among carboxylic acid derivatives. Had you been asked in Chapter 1 to describe the geometry of an amide bond, you probably would have predicted bond angles of 120° about the carbonyl carbon and 109.5° about a tetrahedral amide nitrogen. In the late 1930s, one of Linus Pauling’s most important discoveries was that the geometry of the nitrogen atom of an amide bond in proteins is actually trigonal planar (sp2), not tetrahedral (sp3). A large number of subsequent experiments have verified that amides should be thought of as being best represented as the resonance hybrid of three resonance contributing structures, not two like other carbonyl species. In the unique third contributing structure of an amide, the lone pair on nitrogen donates electron density to create a pi bond between carbon and nitrogen.
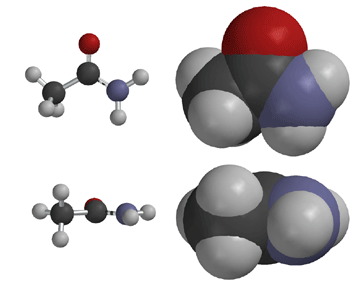
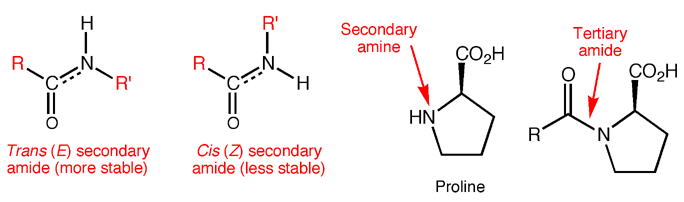
Proteins are made from amino acids connected through mostly secondary amide bonds, which adopt the more stable trans (E) configuration. This is thought to be due to repulsive steric interactions between side chains in the cis (Z) form that are not present in the trans (E) form, along with more favorable dipole-dipole interactions in the trans (E) form. Proline, the only amino acid to contain a secondary amine, produces tertiary amide bonds in proteins, so the cis and trans forms are of similar energy. In fact, proline cis-trans isomerism is now known to contribute substantially to the structure and function of certain proteins.
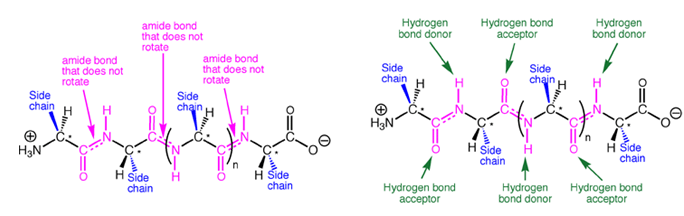
Because of the large rotation barrier of the amide C-N bond, one out of every three bonds in a protein backbone does not rotate, significantly reducing the flexibility of the entire chain. In the context of protein folding, this conformational restriction is essential, because it requires much less stabilization energy to stably fold a chain with conformational restriction every third bond compared to a chain with no bond restrictions. The large number of hydrogen bonds possible along a protein backbone provide directional, non-covalent interactions that work in concert with other non-covalent interactions to stabilize a folded structure.
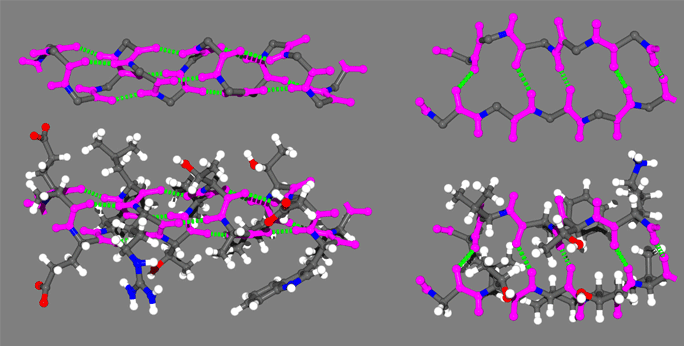
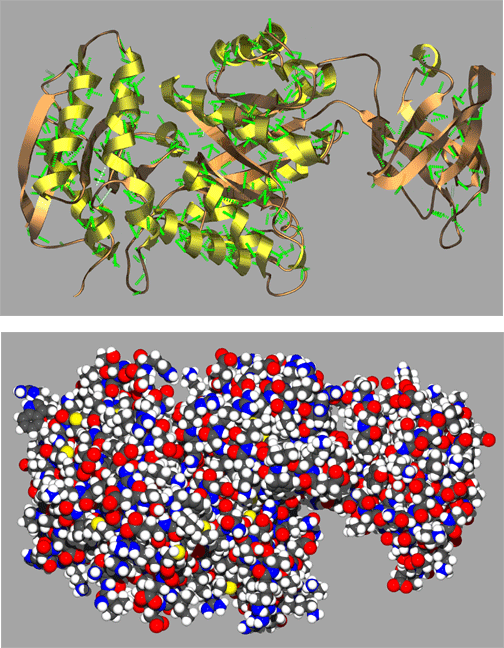
In the above figure are representative protein alpha helix (left) and beta strand (right). The four principle atoms of the backbone amide bonds are highlighted in magenta and backbone hydrogen bonds are shown as broken green lines. The upper structures are the backbone only, with side chain atoms removed for clarity.
A representative complete protein structures in Figure Y illustrates just a
small sample of the remarkably complex and often quite beautiful folding
patterns found in proteins. Here the protein is shown in both a spacefilling
and ribbon cartoon model with coils representing alpha helices and the arrows
representing strands of a beta sheet. Scientists often use visual aids such
as these ribbon cartoons to help them visualize complex structures. Proetin
structures such as these would not be stable if it were not for the hydrogen
bonding and conformational restriction imparted by amide bonds of the protein
backbone. It is reasonable to say that the unique structural characteristics
of amide bonds have played a relatively large part in shaping the evolution
of life on our planet!